Running Experiments
Objectives
Run the training pipeline from the command line
Understand the different execution modes
Use tmux and tee for long-running experiments with logging
Interpret training output: tqdm progress bars, dual losses, early stopping
Understand the two ablation studies (CLI architecture comparison vs notebook hyperparameter sweep)
Navigate the 11-part interactive notebook
Why This Matters
The Scenario: An ML engineer has trained a voltage prediction model that scores 5 mV error on validation data — impressive on paper. But when the hydrogen plant starts operating at 40 bar (a pressure never seen in training), predictions are off by 50 mV. The model memorized the training distribution instead of learning the underlying physics. She needs a training pipeline that proves the model generalizes before deployment.
The Research Question: How do you train and validate 6 different model architectures on the same data with a fair comparison, and how do you use keep-out conditions to measure real generalization rather than just in-distribution accuracy?
What This Episode Gives You: The complete training pipeline — CLI commands, notebook walkthrough, ablation studies, and how to interpret the OOD metrics that actually matter.
Usage
1. Activate the Environment
cd ~/wp7-UC2-pem-electrolyzer-digital-twin
source venv/bin/activate
2. Run the Main Training Script
python scripts/pem_electrolyzer/main.py \
--mode full \
--device cuda \
--epochs 100
3. Interactive Exploration with Jupyter
jupyter lab --no-browser --ip=127.0.0.1 --port=8888
# Open demonstrator-v1.orchestrator.ipynb
Command Line Options
Option |
Default |
Description |
|---|---|---|
|
|
Execution mode: |
|
|
Path to dataset directory |
|
|
Path to output directory |
|
|
Training epochs |
|
|
Random seed |
|
|
Distillation alpha (0.1 = 90% teacher, 10% data) |
|
|
Batch size |
|
|
Learning rate |
|
|
Device: |
Example Experiments
Quick Test (Recommended First)
Run 5 epochs to verify everything works:
python scripts/pem_electrolyzer/main.py --mode quick-test
Full Training Pipeline
Train teacher, distill student, train baselines, and run all evaluations:
python scripts/pem_electrolyzer/main.py --mode full --device cuda --epochs 100
Teacher Only
Train just the physics-hybrid teacher model:
python scripts/pem_electrolyzer/main.py --mode teacher-only --epochs 50 --seed 42
Ablation Study
Run all 7 ablation experiments across 3 seeds (21 total runs):
python scripts/pem_electrolyzer/main.py --mode ablation --device cuda
Custom Learning Rate and Batch Size
python scripts/pem_electrolyzer/main.py --mode full --lr 0.001 --batch-size 2048 --epochs 200
CPU-Only Training
python scripts/pem_electrolyzer/main.py --mode full --device cpu --epochs 50
Reproducibility with Fixed Seed
python scripts/pem_electrolyzer/main.py --mode full --seed 123 --epochs 100
Background Training (Long Runs)
For long-running experiments, use tmux with tee for simultaneous terminal output and logging:
# Start background training with logging
tmux new -s training 'python scripts/pem_electrolyzer/main.py \
--mode full --device cuda --epochs 100 2>&1 | tee training.log'
# Monitor progress from another terminal
tail -f training.log
# Attach to the tmux session
tmux attach -t training
# Detach from session: Ctrl+B, then D
The 2>&1 | tee training.log construct captures both stdout and stderr to the log file while still displaying output in the terminal. This is useful for:
Reviewing training progress after disconnection
Debugging errors that occurred during training
Sharing training logs with collaborators
Expected Training Output
Training uses tqdm progress bars for clean output. When running --mode full, you should see:
=== PEM Electrolyzer PINN Training ===
Device: cuda (NVIDIA A100)
============================================================
Loading Test4 Training Data (STRICT + KEEP-OUT split)
============================================================
Raw samples: 193,248
After strict filtering: 15,234 (7.9%)
KEEP-OUT validation split (76-80C):
Training: 11,534 samples (75.7%)
Validation: 3,700 samples (24.3%)
--- Training Teacher (HybridPhysicsMLP) ---
Teacher: |████████████████████| 100/100 [01:02<00:00] loss=0.00032 val=13.9mV best=13.9mV lr=0.00010 *
--- Training Baselines ---
Training pure_mlp (2,049 params)...
Training big_mlp (43,393 params)...
Training transformer (529,793 params)...
--- Distilling Student (PhysicsHybrid12Param) ---
Student: |███████████████ | 152/200 [01:18<00:24] Early stop @ 152 | best=13.8mV
--- OOD Evaluation (All 6 Models) ---
FULL MODEL COMPARISON (All 6 Models)
==========================================================================
Model Params Val (mV) Test2 (mV) Test3 (mV) OOD Avg (mV)
--------------------------------------------------------------------------
Distilled Student 12 13.8 14.7 15.9 15.3
Teacher 9354 13.9 42.5 13.8 28.2
Pure Physics 12 25.1 18.3 22.7 20.5
PureMLP 2049 12.5 45.2 38.1 41.7
BigMLP 43393 11.8 52.3 41.5 46.9
Transformer 529793 10.2 68.4 55.2 61.8
Results saved to results/results.json
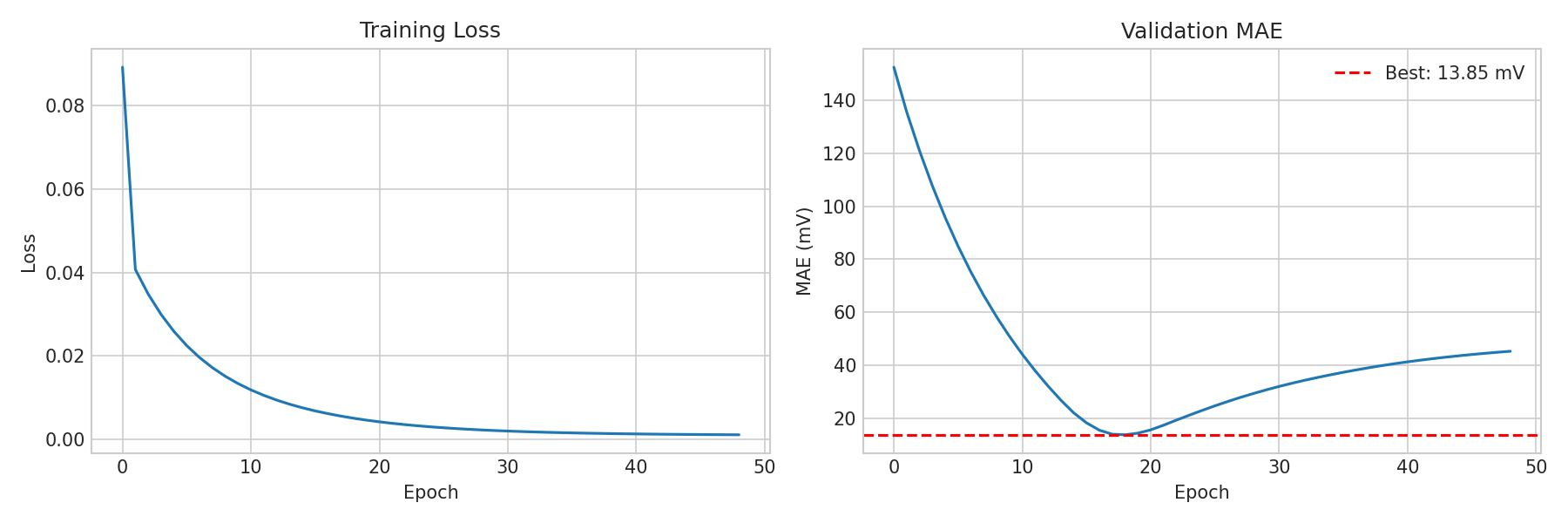
Real training curves from the teacher model: (Left) MSE loss decreasing smoothly over epochs. (Right) Validation MAE converging to 13.85 mV best, with the dashed red line marking the best checkpoint.
Key observations from the output:
Teacher converges early (typically best at epoch ~17) with early stopping patience of 30
Student training shows label loss (L) and distillation loss (D) separately
Progress bars use tqdm with real-time postfix: loss, validation MAE, best MAE, learning rate
A
*after the postfix indicates a new best epoch was foundThe 6-model comparison table shows all models sorted by OOD Average MAE
The distilled student (12 params) typically beats all larger models on OOD
If you see CUDA not available, falling back to CPU, GPU drivers may not be installed correctly. See Episode 03 for troubleshooting.
Training Deep Dive
What Happens During --mode full
The full pipeline executes these steps in order:
Load and filter data: Load Test4 CSV, apply strict filtering (removes out-of-range samples), split into train/validation using 76-80C keep-out
Train teacher (HybridPhysicsMLP): SGD optimizer, CosineAnnealingLR, 100 epochs with early stopping (patience=30)
Train baselines: PureMLP, BigMLP, Transformer — each with the same SGD config for fair comparison
Distill student (PhysicsHybrid12Param): Adam optimizer, ReduceLROnPlateau, 200 epochs (patience=40), alpha=0.1
Train pure physics: Same student architecture but alpha=1.0 (no teacher signal)
OOD evaluation: Evaluate all 6 models on Test2 and Test3 datasets
Reading the Progress Bar
The tqdm progress bar shows real-time training metrics:
Teacher: |████████████████████| 100/100 [01:02<00:00] loss=0.00032 val=13.9mV best=13.9mV lr=0.00010 *
Field |
Meaning |
|---|---|
|
Training MSE loss (in Volts squared). Lower is better. |
|
Current epoch’s validation MAE (in millivolts) |
|
Best validation MAE seen so far across all epochs |
|
Current learning rate (decreases via scheduler) |
|
This epoch achieved a new best validation MAE |
For the student, the progress bar also shows dual losses:
Student: |███████████████ | 152/200 [01:18<00:24] L=0.00045 D=0.00032 val=13.8mV best=13.8mV
Field |
Meaning |
|---|---|
|
Label loss: MSE(student predictions, true labels) |
|
Distillation loss: MSE(student predictions, teacher predictions) |
With alpha=0.1, the total loss is 0.1 x L + 0.9 x D. The distillation loss (D) typically dominates.
Early Stopping Behavior
Training does not always run for the full epoch count:
Teacher: Typically converges at epoch ~17-30 (patience=30 means it runs 30 more epochs after the best, then stops)
Student: May early-stop around epoch 100-160 (patience=40)
Baselines: Variable — simpler models converge faster, Transformer may use more epochs
When early stopping triggers, the progress bar shows: Early stop @ 152 | best=13.8mV
The best model checkpoint is saved at the epoch with the lowest validation MAE, not the final epoch.
Learning Rate Schedules
Two different schedules are used:
CosineAnnealingLR (teacher and baselines): Smoothly decreases the learning rate from 0.01 to 0.0001 following a cosine curve over the full epoch count. This provides gradual refinement.
ReduceLROnPlateau (student): Starts at 0.001 and halves the learning rate whenever validation MAE does not improve for 10 consecutive epochs. This adapts to the student’s convergence speed.
Ablation Studies
There are two distinct ablation studies — one for the CLI and one for the notebook. They test different hypotheses.
CLI Ablation (--mode ablation)
Compares model architectures to prove physics-informed models generalize better than pure ML:
python scripts/pem_electrolyzer/main.py --mode ablation --device cuda
# |
Experiment |
What It Tests |
|---|---|---|
1 |
|
Small MLP baseline (~2K params, no physics) |
2 |
|
Large MLP baseline (~43K params, no physics) |
3 |
|
Attention-based baseline (~530K params, no physics) |
4 |
|
Physics-hybrid teacher (~9K params) |
5 |
|
Student with alpha=1.0 (12 params, no teacher signal) |
6 |
|
Student with alpha=0.1, keep-out validation |
7 |
|
Student with alpha=0.1, random validation |
Each experiment runs across 3 seeds [42, 91, 142] = 21 total runs. Results are saved to results/ablation/ with per-experiment JSON files.
Key question answered: Does physics help, or can a large enough network learn it from data?
Notebook Ablation (Part 9)
Compares hyperparameter choices to show which training decisions matter most:
# |
Experiment |
What It Tests |
|---|---|---|
1 |
|
Default config: SGD + keep-out + lr=0.01 |
2 |
|
Temperature keep-out disabled |
3 |
|
80/20 random validation split |
4 |
|
Adam optimizer instead of SGD |
5 |
|
Learning rate 0.001 (10x lower) |
6 |
|
Learning rate 0.05 (5x higher) |
7 |
|
No weight decay (weight_decay=0) |
Each experiment runs across 3 seeds [42, 123, 456] = 21 total runs. Disabled by default — set RUN_ABLATION = True in the notebook to run.
Key question answered: Which training choices matter most for OOD generalization?
Expected Ablation Insights
The paper quantifies three complementary mechanisms that contribute to the student’s OOD performance:
Mechanism |
Improvement |
Comparison |
|---|---|---|
Physics constraints |
56% |
Pure MLP (35.1 mV) → Student (15.3 mV) |
Knowledge distillation |
50% |
Pure Physics (30.6 mV) → Student (15.3 mV) |
Keep-out validation |
29% |
Random val (22.3 mV) → Keep-out (15.9 mV) |
Additional insights:
SGD beats Adam for the teacher by ~10-15 mV on OOD
Weight decay has a modest effect (~2-3 mV)
Learning rate sensitivity is moderate — 0.01 is near-optimal; 0.05 can diverge, 0.001 converges slowly
Architecture matters most: the 12-param student beats ~530K-param baselines regardless of hyperparameters
Reproducibility Across Seeds
Results are stable across random seeds [42, 91, 142]:
Model |
OOD Avg (mean ± std) |
|---|---|
Distilled Student |
15.9 ± 1.0 mV |
Teacher |
28.2 ± 1.5 mV |
Pure MLP |
35.1 ± 2.3 mV |
Transformer |
39.0 ± 3.1 mV |
The worst Student seed (17.1 mV) still beats the best Transformer seed (37.2 mV), confirming that the physics-informed approach is robust to initialization.
Notebook Walkthrough (11 Parts)
The interactive notebook (demonstrator-v1.orchestrator.ipynb) covers the full pipeline in 11 parts:
Part |
What Happens |
Key Output |
|---|---|---|
1 |
Environment setup |
GPU check, imports |
2 |
Data exploration |
Distribution plots, filtering statistics |
3 |
PINN methodology |
Voltage decomposition stacked area chart |
4 |
Teacher training |
tqdm progress bar, training curves, best epoch |
5 |
Baseline comparison |
Train PureMLP, BigMLP, Transformer, Pure Physics; 5-model bar chart |
6 |
Knowledge distillation |
Student training with label/distillation loss breakdown |
7 |
6-model comparison |
Full comparison table + bar chart + residual distribution |
8 |
Interactive widgets |
Configure and launch experiments with dropdown controls |
9 |
Ablation study |
Systematic parameter sweeps (optional, set RUN_ABLATION=True) |
10 |
Summary |
Model ranking, next steps |
11 |
12-number equation |
Extract 12 params, NumPy implementation, verification, polarization curves |
Part 11: The Biggest Takeaway
Part 11 is the culmination of the entire notebook. It demonstrates that:
The trained student is defined by exactly 12 scalar constants
A standalone NumPy function reproduces the model with zero PyTorch dependencies
The NumPy function matches PyTorch
model.forward()to micro-volt precisionFull voltage decomposition is computed step-by-step for sample operating conditions
Polarization curves are generated showing temperature and pressure effects
A reproducibility card lists all constants ready to copy-paste
Result File Format
Results are saved to results/results.json with this structure:
Field |
Type |
Description |
|---|---|---|
|
string |
Training timestamp |
|
object |
Training configuration (seed, epochs, lr, alpha, device) |
|
object |
Teacher metrics (val_mae_mV, test2_mae_mV, test3_mae_mV, ood_avg_mV) |
|
object |
Student metrics + learned physics and hybrid parameters |
The student section includes the 12 learned parameters:
{
"student": {
"val_mae_mV": 13.81,
"test2_mae_mV": 14.67,
"test3_mae_mV": 15.91,
"ood_avg_mV": 15.29,
"physics_params": {
"i_lim_ref": 1.293,
"i0_a_ref": 8.075e-05,
"i0_c_ref": 0.003470,
"R_ohm_ref": 0.9947,
"alpha_a": 0.6375,
"alpha_c": 0.6710
},
"hybrid_params": {
"corr_a": 0.03471,
"corr_b": 0.6046,
"corr_c": 0.6341,
"corr_d": 0.02045,
"corr_p": 1.8518,
"corr_t": 0.3209
}
}
}
These 12 numbers are all that is needed to reproduce the model’s predictions without retraining. See Episode 07 for how to use them.
Keypoints
Always run
--mode quick-testfirst to verify setupUse
--device cudafor GPU training (much faster)Use tmux with
teefor long-running experiments with loggingTraining uses tqdm progress bars: watch for
loss,val,best,lr, and*(new best)Student shows dual losses: L (label) and D (distillation), weighted by alpha
Early stopping saves the best checkpoint, not the final epoch
Two ablation studies: CLI compares architectures (7 experiments x 3 seeds), notebook compares hyperparameters (7 x 3)
Keep-out validation is the single most important training choice for OOD
The notebook has 11 parts: Parts 5 and 7 compare all 6 models, Part 9 is ablation, Part 11 is the 12-number equation
Results include 12 learned parameters in
results.jsonfor full reproducibility